Reading Writing And Editing Dna
Revealing RNA's Hidden Dance: A New Frontier in Molecular Imaging
A revolutionary study using Individual-Particle cryo-Electron Tomography (IPET) unveils RNA’s folding dynamics at the single-molecule level, and could pave the way for better RNA vaccines and molecular sensors

[Grok]
RNA is the acrobat of biomolecules, twisting and folding into a multitude of shapes based on its environment. While this structural flexibility is crucial for its biological functions, it also makes RNA notoriously hard to study. Traditional imaging methods, such as cryo-electron microscopy (cryo-EM) single-particle averaging (SPA) analysis, rely on combining data from thousands of molecules with similar shapes. This averaging approach blurs the distinctiveness of individual RNA molecules, leaving researchers with an incomplete picture.
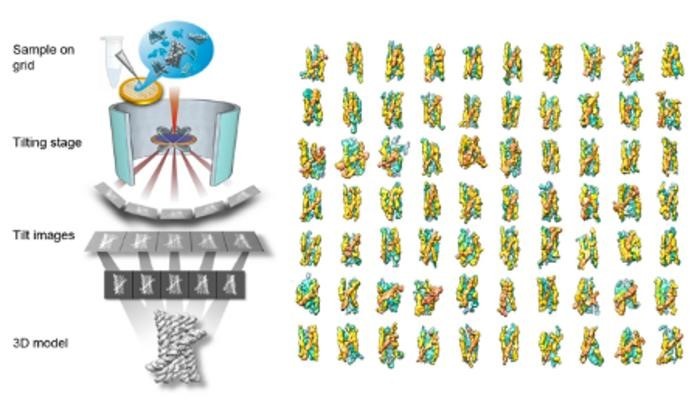
But a groundbreaking study published in Nature Communications changes the game. Scientists from the Molecular Foundry at Lawrence Berkeley National Laboratory and the Interdisciplinary Nanoscience Center at Aarhus University have introduced a new dimension to RNA research: the ability to study the 3D structure of single molecules without averaging. Their tool of choice? Individual-Particle cryo-Electron Tomography (IPET), an advanced imaging technique that homes in on the 3D shapes of cryo-preserved molecules, one particle at a time.
From Impossible to Insightful: The Power of IPET
Until recently, visualizing single-molecule 3D structures seemed like a pipe dream. The weak signals generated by individual particles rendered this approach ineffective. “It was considered a dead-end method since the 1970s,” recalls Gang Ren, a staff scientist at the Molecular Foundry and co-leader of the study with Ebbe Andersen from Aarhus University. However, technological advancements have breathed new life into this method, transforming it into a powerful tool for molecular exploration.
In this study, the researchers applied IPET to RNA origami—engineered RNA molecules designed to fold into precise nanoscale structures. While previous studies using cryo-EM SPA provided static snapshots of RNA origami’s 3D structure, they failed to unravel its folding dynamics. IPET, however, illuminated the entire folding journey. By capturing RNA molecules in various stages of folding—from immature configurations to their optimal forms—the researchers essentially created a "movie" of RNA’s dynamic transformation.
Capturing RNA’s Dynamic Folding
This molecular movie revealed critical insights. For instance, the team observed a folding trap, a point where RNA temporarily gets stuck, before transitioning to a more compact and functional shape. Such observations not only deepen our understanding of RNA’s folding process but also hold immense promise for practical applications.
“The IPET technique provides us with a more dynamic view of the molecular world,” explains Andersen. “It is our hope that this insight will enable us to engineer the folding of more effective RNA vaccines and dynamic sensors for molecular medicine.”
A video illustrating this folding process can be viewed below. The animation demonstrates how IPET zooms in on a single molecule, revealing its intricate 3D structure and capturing the fluidity of RNA’s folding landscape.
Behind the Scenes: IPET and Cryo-EM SPA
What is IPET?
Individual-Particle Cryo-Electron Tomography (IPET) is a revolutionary technique for imaging single macromolecular particles in 3D at low to intermediate resolution. Unlike cryo-EM SPA, it doesn’t rely on averaging images from thousands of molecules. Instead, it combines innovations like cryo-electron tomography, advanced reconstruction algorithms, missing-wedge correction, contrast enhancement, and graphene grids to visualize individual molecules in their natural state.
Cryo-EM SPA: The Traditional Approach
Cryo-EM SPA has been a gold standard for high-resolution imaging of biomolecules. It freezes molecules in a glassy ice state, capturing detailed 3D reconstructions by averaging thousands of similar particles. However, this method falls short when studying dynamic molecules like RNA that continuously change shape.
Looking Ahead: A New Era for RNA Research
The ability to capture RNA’s ever-shifting forms could revolutionize fields ranging from vaccine development to molecular medicine. By peeling back the layers of RNA’s dynamic folding process, IPET has set a new standard for single-molecule imaging, proving that what was once deemed impossible is now within reach.
Read More
Newletter & More

SynBioBeta
Join the innovators shaping the future with SynBio + AI. From health to ag, materials & more—be part of the revolution.
















